Porcine Endogenous Retroviruses: Advancements in Safe Pig-to-Human Organ Xenotransplantation
Written and cover illustrated by Laurianne Daoust.
The demand for organ transplants is at an all-time high. In 2022, over 42,000 organ transplants were performed in the United States of America [1]. However, the limited supply of organ donations is a pressing, ongoing issue. Today, the US national transplant waiting list counts over 103,000 people and, everyday, about 17 people die while waiting for an organ transplant [1].
Xenografts—organs or tissues donated from one species to another—are a viable solution for the organ shortage [2, 3]. Pig xenografts are particularly promising due to their similarities to human organs in size and function, short breeding time, and large litters [2, 3]. Despite these advantages, porcine organs pose a risk of infection to human recipients as porcine endogenous retroviruses (PERVs) are integrated into the pig genome and can insert their own genetic material in the human host genome [3]. Moreover, genes that encode for PERVs are scattered throughout the pig genome, making them notably difficult to eliminate [3]. Thankfully, researchers have come up with two promising solutions to prevent PERV transmission to human recipients: 1) the production of transgenic pigs expressing PERV siRNAs, and 2) genetic editing of the pig genome through CRISPR-Cas9 technology.
1) Production of Transgenic Pigs Expressing PERV siRNAs
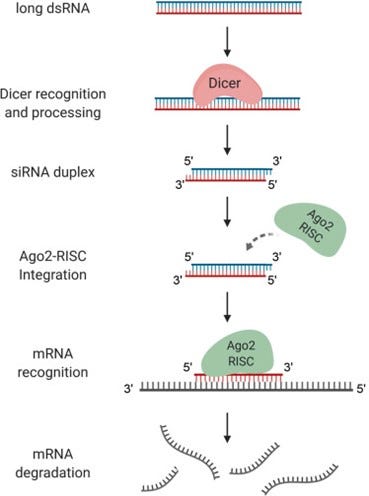
Developed in the early 2000s, RNA interference (RNAi) silences genes by blocking their translation from DNA into protein [4-6]. The process is initiated by the ribonuclease III enzyme Dicer, which cleaves sequence-specific target mRNAs into short double-stranded RNA segments (see Figure 1) [4, 7]. These segments are known as small interfering RNAs (siRNAs), and they assemble with the RNA-induced silencing complex (RISC) in the cytoplasm. This complex specifically recognizes target mRNA sequences homologous to the siRNA. Nucleases (here, the Ago2 protein) bind to and degrade the mRNA, preventing protein translation and subsequent expression of the target gene [4, 8].
Figure 1. Schematic of the RNAi mechanism [9].
By using PERV-specific siRNAs, researchers were able to reduce PERV expression in vitro [4, 10]. In fact, using synthetic siRNAs designed to target various segments of the PERV viral genes—specifically gag, pol, and env—Karlas et al. successfully identified potential RNAi target sites within the PERV genome [4]. Among these, the pol2 sequence—encoding for reverse transcriptase—has the highest efficacy in reducing virus expression. This RNAi strategy raises the prospect of generating transgenic pigs providing safe xenotransplants. However, no study has shown complete suppression of PERV expression using siRNAs so far [4, 10].
2) Genetic Editing of the Pig Genome Using CRISPR-Cas9
Genome editing is an emerging technology which inactivates single genes in cells [11, 12]. A main breakthrough in genetic editing involves Clustered Regularly Interspaced Short Palindromic Repeats—or CRISPR—and associated protein Cas9.
The CRISPR-Cas9 system was inspired by bacteria’s ability to recognize foreign genetic elements from viral infections [13]. When infected, the bacteria integrate sections of the viral genome into their own, forming CRISPR DNA arrays. These enable bacteria to "remember" previous invaders and attack them upon reinfection. Bacteria produce RNA segments from CRISPR arrays, which recognize and attach to viral DNA regions.
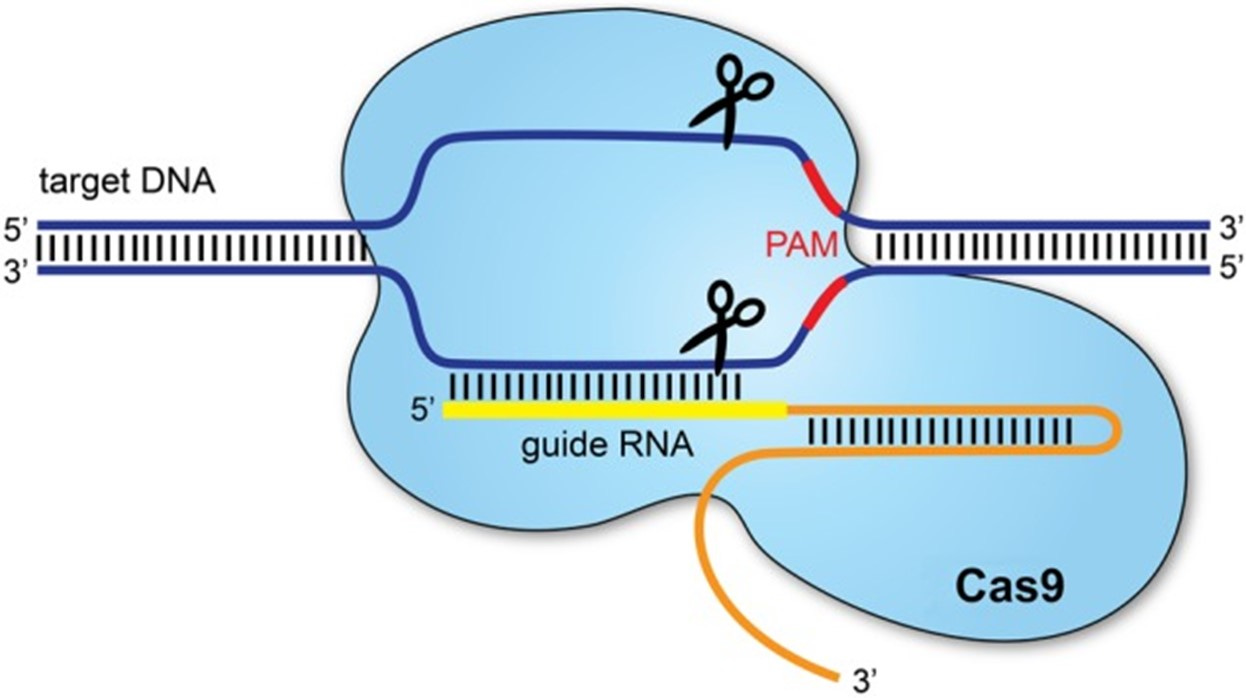
As shown in Figure 2, Cas9 acts as molecular scissors to cleave invader DNA, causing a double-stranded break [12, 13]. A protospacer adjacent motif (PAM) sequence aids the binding of Cas9. Researchers have combined this immune defense mechanism with guide RNA (gRNA), a small piece of RNA with a guide sequence which binds to a target sequence. When bound to Cas9, the system cuts in precise regions to knock-out carefully chosen target genes [13]. As an attempt to repair the cut, the cell will either insert or delete fragments of DNA (i.e., non-homologous end joining), disrupting gene functionality, or utilizes a custom template to fix itself (i.e., homology directed repair) [13].
Figure 2. The CRISPR/Cas9 system [13].
By altering the gRNA sequence, CRISPR-Cas9 can precisely target the catalytic region of the pol gene, which encodes for enzymes crucial to retrovirus replication (i.e., reverse transcriptase, protease, and integrase) [14, 15]. Knocking out this gene halts PERV replication, preventing the infection of human recipients.
Using CRISPR-Cas9 technology, Yang et al. demonstrated a 1000-fold reduction in PERV infectivity, eradicating all copies of PERVs in porcine kidney epithelial cells [16]. Moreover, the scientists created PERV-free piglets by inactivating all PERV copies in primary fibroblast cells [14, 15]. Therefore, CRISPR-Cas9 ensures complete and precise elimination of PERVs [16].
However, CRISPR-Cas9 faces limitations, including off-target effects—when Cas9 cleaves untargeted genomic sites—and mosaicism—when CRISPR-Cas9 does not uniformly edit every cell, leading to the coexistence of edited and unedited cells [17, 18]. This can cause adverse effects in human recipients and necessitates ongoing monitoring. Regarding long-term viability, once retroviral sequences are knocked out in pigs, their piglets inherit PERV-inactivated genes. [14]. However, inbreeding of the genetically modified animals when breeding pigs for xenotransplantation in abundance is a main obstacle [2]. Nonetheless, CRISPR-Cas9 is the most promising method of preventing PERV transmission in xenotransplantation thanks to its high effectiveness in PERV elimination and heritability.
In response to the growing organ transplant demand, the PERV mitigation becomes critical in pig-to-human xenotransplantation. Although the production of transgenic pigs expressing PERV siRNAs reduces risks of adverse effects by avoiding direct genetic modification, this method requires constant monitoring as it does not completely eliminate PERV expression. On the other hand, CRISPR-Cas9 allows for high precision targeting and permanent elimination of PERVs, ensuring the safety of human recipients. With further research, we could limit the risks of off-target mutations and mosaicism, potentially solving the global organ shortage.
References
HRSA. "Organ Donation Statistics." https://www.organdonor.gov/learn/organ-donation-statistics.
J. Denner, "Porcine Endogenous Retroviruses and Xenotransplantation, 2021," Viruses, vol. 13, no. 11, Oct 26 2021, https://doi.org/10.3390/v13112156.
J. Y. C. Yang and M. M. Sarwal, "Transplant genetics and genomics," Nature Reviews Genetics, vol. 18, no. 5, pp. 309-326, 2017, https://doi.org/10.1038/nrg.2017.12.
A. Karlas, R. Kurth, and J. Denner, "Inhibition of porcine endogenous retroviruses by RNA interference: increasing the safety of xenotransplantation," Virology, vol. 325, no. 1, pp. 18-23, 2004, https://doi.org/10.1016/j.virol.2004.04.022.
N. J. Caplen, S. Parrish, F. Imani, A. Fire, and R. A. Morgan, "Specific inhibition of gene expression by small double-stranded RNAs in invertebrate and vertebrate systems," Proceedings of the National Academy of Sciences, vol. 98, no. 17, pp. 9742-9747, 2001, https://doi.org/10.1073/pnas.171251798.
P. Guo et al., "Engineering RNA for targeted siRNA delivery and medical application," Adv Drug Deliv Rev, vol. 62, no. 6, pp. 650-66, 2010, https://doi.org/10.1016/j.addr.2010.03.008.
S. M. Hammond, E. Bernstein, D. Beach, and G. J. Hannon, "An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells," Nature, vol. 404, no. 6775, pp. 293-296, 2000, doi: https://doi.org/10.1038/35005107.
E. Billy, V. Brondani, H. Zhang, U. Müller, and W. Filipowicz, "Specific interference with gene expression induced by long, double-stranded RNA in mouse embryonal teratocarcinoma cell lines," Proceedings of the National Academy of Sciences, vol. 98, no. 25, pp. 14428-14433, 2001, https://doi.org/10.1073/pnas.261562698.
J. Cortinhas, A. P. Pêgo, and P. M. D. Moreno, "Chapter 9 - Toward the therapeutic application of small interfering RNA bioconjugates in the central nervous system," in Handbook of Innovations in Central Nervous System Regenerative Medicine, A. J. Salgado Ed.: Elsevier, 2020, pp. 333-374.
S. Miyagawa et al., "Prevention of PERV Infections in Pig to Human Xenotransplantation by the RNA Interference Silences Gene," The Journal of Biochemistry, vol. 137, no. 4, pp. 503-508, 2005, https://doi.org/10.1093/jb/mvi059.
F. Tanihara, M. Hirata, and T. Otoi, "Current status of the application of gene editing in pigs," Journal of Reproduction and Development, vol. 67, no. 3, pp. 177-187, 2021, https://doi.org/10.1262/jrd.2021-025.
C. Erlbaum, "Xenotransplantation: The Science, the Advantages, the Ethics.," Touro College, vol. 12, no. 1, pp. 53-58, 2018. [Online]. Available: https://touroscholar.touro.edu/sjlcas/vol12/iss1/9/.
R. Melody, K. Andrew, W. Caroline, and K. David, "What is CRISPR/Cas9?," BMJ, vol. 101, no. 4, p. 213, 2016, https://doi.org/10.1136/archdischild-2016-310459.
D. Niu et al., "Inactivation of porcine endogenous retrovirus in pigs using CRISPR-Cas9," Science, vol. 357, no. 6357, pp. 1303-1307, 2017, https://doi.org/10.1126/science.aan4187.
L. Yang et al., "Genome-wide inactivation of porcine endogenous retroviruses (PERVs)," Science, vol. 350, no. 6264, pp. 1101-1104, 2015, https://doi.org/10.1126/science.aad1191.
D. Niu et al., "Porcine genome engineering for xenotransplantation," Advanced Drug Delivery Reviews, vol. 168, pp. 229-245, 2021, https://doi.org/10.1016/j.addr.2020.04.001.
N. Ryczek, M. Hryhorowicz, J. Zeyland, D. Lipiński, and R. Słomski, "CRISPR/Cas Technology in Pig-to-Human Xenotransplantation Research," Int J Mol Sci, vol. 22, no. 6, 2021, https://doi.org/10.3390/ijms22063196.
M. Mehravar, A. Shirazi, M. Nazari, and M. Banan, "Mosaicism in CRISPR/Cas9-mediated genome editing," Developmental Biology, vol. 445, no. 2, pp. 156-162, 2019, https://doi.org/10.1016/j.ydbio.2018.10.008.



Super clear and well written. Excited to see where these advances go!